Bits of the Batch


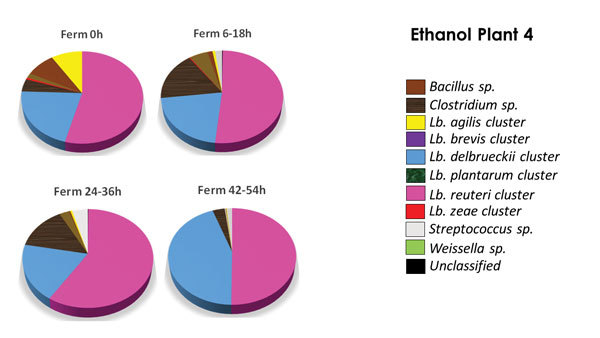

PHOTO: LALLEMAND BIOFUELS & DISTILLED SPIRITS
March 27, 2020
BY Lisa Gibson
Advertisement
Advertisement
Related Stories
A pan-European road trip featuring cars and trucks powered by renewable fuels logged 77,500 kilometers across 16 countries in a real-world demonstration that these fuels deliver significant GHG reductions.
International Air Transport Association has announced the release of the Sustainable Aviation Fuel (SAF) Matchmaker platform, to facilitate SAF procurement between airlines and SAF producers by matching requests for SAF supply with offers.
ATR and French SAF aggregator ATOBA Energy on June 19 signed a memorandum of understanding (MOU) to explore ways to facilitate and accelerate sustainable aviation fuel (SAF) adoption for ATR operators.
The USGC has added Sarah McLeeson as its new trade policy coordinator. McLeeson will organize staff and member travel, make arrangements for visiting trade teams and facilitate program planning and execution.
International Sustainability & Carbon Certification has announced that Environment and Climate Change Canada has approved ISCC as a certification scheme in line with its sustainability criteria under its Clean Fuel Regulations.





